COVID-19 AND SARS-CoV-2 Research
2020
Pathogenic Priming Likely Contributes to Serious and Critical Illness and Mortality in COVID-19 via Autoimmunity -
Pathogenic Priming Likely Contributes to Serious and Critical Illness and Mortality in COVID-19 via Autoimmunity James Lyons-Weiler PII: S2589-9090(20)30018-6 DOI: https://doi.org/10.1016/j.jtauto.2020.100051 Reference: JTAUTO 100051 To appear in: Journal of Translational Autoimmunity Received Date: 9 March 2020 Revised Date: 31 March 2020 Accepted Date: 2 April 2020 Please cite this article as: J. Lyons-Weiler, Pathogenic Priming Likely Contributes to Serious and Critical Illness and Mortality in COVID-19 via Autoimmunity, Journal of Translational Autoimmunity, https:// doi.org/10.1016/j.jtauto.2020.100051.
Abstract Homology between human and viral proteins is an established factor in viral- or vaccine-induced autoimmunity. Failure of the SARS and MERS vaccines in animal trials involved pathogenesis consistent with an immunological priming that could involve autoimmunity in lung tissues due to previous exposure to the SARS and MERS spike protein. Exposure to SARS-CoV-2 pathogenesis in COVID-19 likely will lead to similar outcomes. Immunogenic peptides in viruses or bacteria that match human proteins are good candidates for pathogenic priming peptides (similar to the more diffuse idea of “immune enhancement”). Here I provide an assessment of potential for human pathogenesis via autoimmunity by exposure, through infection or injection, SAR-CoV-2 spike proteins, and all other SARS-CoV-2 proteins. Immunogenic epitopes in each SARS-CoV-2 protein were compared to human proteins in search of high local homologous matching. Only one immunogenic epitope in a SARS-CoV-2 had no homology to human proteins. Mapping of the genes encoding human protein matches to pathways point to targets that could explain the observed presentation of symptoms in COVID-19 disease. It also strongly points especially to surprisingly large number of opportunities for expected disturbances in the immune system itself, targeting elements of MHC Class I and Class II antigen presentation, PD-1 signaling, cross-presentation of soluble exogenous antigens and the ER-Phagosome pathway. Translational consequences of these findings are explored. [Donate]
#2 We have found a protein motif "signature" of pathogenicity that seems characteristic of 2019-nCoV. Specifically, the oldest sequence we have to date (from 2005) that shares the motif fingerprint is HK-3. HK-1 and HK-2 share part of the pathogenicity signature, but only HK-3 has the complete match. Research Report here.
IPAK RESEARCH REPORT COVID-19-2020_1
"Motif Pathogenicity Typing: SARS-CoV-2 Coronavirus May Contain a Unique “Likely Pathogenic” Protein Motif Signature Also Found in a Natural Isolate from 2005" Download_PDF
Abstract
Careful analysis of protein motif signatures and iterative phylogenetic analysis of amino acid sequences of coronaviruses reveal a characteristic functional motif signature of the coronavirus first detected in afflicted humans in China in December, 2019, currently responsible for the deaths of over 700 people so far and massive isolation efforts in China. New results show that original conjectures the modified protein signatures as indicative of supporting a likely laboratory origin via recombination technology do not, at this time, appear to be not supported. New data indicate that invoking a recent laboratory intentional modification is not necessary to understand the origins of the pathogenicity. This, however, does not rule out a laboratory escape of pathogenic virus for pathogenicity or vaccine development research. Continuous ongoing detailed sequence analysis is necessary. An evolutionary awareness of the inheritance and acquisition of characteristic functional motif signatures will aid in understanding limits of applicability such studies; studies of “SARS” and “SARS-Like” coronaviruses cannot be expected to be useful to generalize about the pathogenicity from sequence or structural insights alone. Determining, recording and publishing functional motif signatures could prove essential for communicating pathogenic-like coronavirus types with respect to expected pathogenicity. Laboratories with natural isolates, isolates from humans, and recombined coronaviruses that include spike protein sequences should analyze the spike protein for the putatively characteristic pathogenic functional motif signature: which appears at this time to include a shortened N-terminal spike domain, missing She-3 and KxDL motifs in the Spike 2 segment, and a C-terminal Gp41 (retroviral envelope) motif. Research into C-terminal motifs and elements may prove to potentially useful for tracking laboratory modified coronavirus types. The presence of the proposed pathogenicity signature and an understanding of the provenance of the sequence information involves laboratory origin of related sequences, but seems at this time to rule out recent laboratory origin of the SARS-CoV-2 lineage. A recently reported furin-like cleavage site that has a similar phylogenetic distribution in B-coronaviruses also seems like a promising lead for therapeutics.
Notice
IPAK is at the leading edge of understanding where the SARS-CoV-2 Coronavirus originated, and while we do not have (and at this time, 2/7/2020), and no one else has a solid, definitive answer, our analyses have been instrumental in asking tough questions about the origins of SARS-CoV-2.
In January, we analyzed sequences to determine if a link exists between wild bat coronaviruses and lab-housed viruses, and we found a possible connection to a vector sequence technology that originated in China called pShuttle-SN. Upon further analysis, the link is clearly not a direct one; however, the result was not spurious. The original published analysis of the 2019-nCoV genome had a segment, later referred to as "the middle fragment", that no one could match to any known sequences. Our analysis pointed to pShuttle-SN, and thereby to spike proteins.
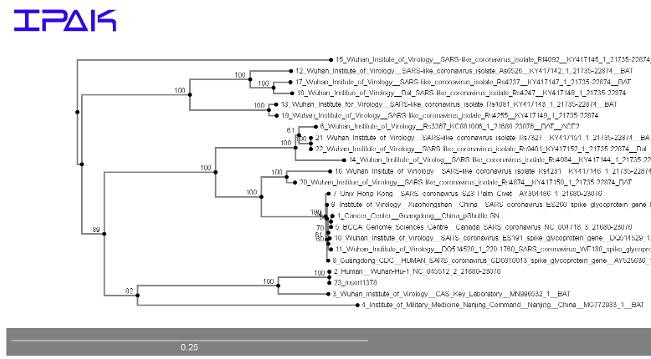
Updates and new data still support a potential link to laboratory-housed sequences in Nanjiang and in Wuhan as the most similar relatives given data from the Spike protein in 2019-nCoV. Our results, summarized below, are preliminary, but are based on standard phylogenetic techniques (description)
We call upon Scientists in all laboratories in China and all countries to sequence all housed, propagated, and stored samples of coronaviruses. New data are necessary to flesh out these relationships both with whole genome, multi-gene, single-gene and nucleotide base-level analyses. We are calling upon data analysts to study these sequences, especially with respect to the genomic location and protein motif locations,and subsequences around in withtin the coronavirus Spike protein coding genes in published sequences for evidence of recombination nearby, and within, the gene.
To send news about new sequence data, and new results using existing coronavirus data, to IPAK, contact info@ipaknowledge.org or tweet a link to @lifebiomedguru. We will share all reported results on our social media outlets.
To support IPAK's research in molecular epidemiology and the origins of the 2019-nCoV coronavirus, please visit our "How To Donate" page. Small monthly donations will power these analyses. We will be submitting our full analyses for peer-reviewed publication when sufficient data are available. IPAK is a registered not-for-profit corporation in the Commonwealth of Pennsylvania, but is not yet a 501c3. Your donation is not tax deductible at this time. [Donate]